Introduction¶
torch-sla (Torch Sparse Linear Algebra) is a memory-efficient library for sparse linear algebra in PyTorch. It provides differentiable sparse linear equation solvers with multiple backends, supporting both CPU and CUDA.
Key Features¶
- Memory Efficient: Only stores non-zero elements — solve systems with millions of unknowns using minimal memory
- Multiple Backends: Choose from SciPy, Eigen (C++), cuDSS, or PyTorch-native
- Backend/Method Separation: Independently specify the backend and solver method
- Auto-selection: Automatically choose the best backend and method based on device, dtype, and problem size
- Gradient Support: Full gradient computation via PyTorch autograd with O(1) graph nodes
- Batched Operations: Support for batched sparse tensors with shape
[..., M, N, ...] - Property Detection: Automatic detection of symmetry and positive definiteness
- Distributed Support: Distributed sparse matrices with halo exchange for parallel computing
- Large Scale: Tested up to 169 million DOF with near-linear scaling
Recommended Backends¶
Based on extensive benchmarks on 2D Poisson equations (tested up to 169M DOF):
Problem Size |
CPU |
CUDA |
Notes |
|---|---|---|---|
Small (< 100K DOF) |
|
|
Direct solvers, machine precision |
Medium (100K - 2M DOF) |
|
|
cuDSS is fastest on GPU |
Large (2M - 169M DOF) |
N/A |
|
Iterative only, ~1e-6 precision |
Very Large (> 169M DOF) |
N/A |
|
Multi-GPU domain decomposition |
Key Insights¶
PyTorch CG+Jacobi scales to 169M+ DOF with near-linear O(n^1.1) complexity
Direct solvers limited to ~2M DOF due to memory (O(n^1.5) fill-in)
Use float64 for best convergence with iterative solvers
Trade-off: Direct = machine precision (~1e-14), Iterative = ~1e-6 but 100x faster
Core Classes¶
SparseTensor¶
The main class for sparse matrix operations. Supports batched and block sparse tensors.
from torch_sla import SparseTensor
# Simple 2D matrix [M, N]
A = SparseTensor(values, row, col, (M, N))
# Batched matrices [B, M, N]
A = SparseTensor(values_batch, row, col, (B, M, N))
# Solve, norm, eigenvalues
x = A.solve(b)
norm = A.norm('fro')
eigenvalues, eigenvectors = A.eigsh(k=6)
SparseTensorList¶
A list of SparseTensors with different sparsity patterns.
from torch_sla import SparseTensorList
matrices = SparseTensorList([A1, A2, A3])
x_list = matrices.solve([b1, b2, b3])
DSparseTensor¶
Distributed sparse tensor with domain decomposition and halo exchange.
from torch_sla import DSparseTensor
D = DSparseTensor(val, row, col, shape, num_partitions=4)
x_list = D.solve_all(b_list)
LUFactorization¶
LU factorization for efficient repeated solves with same matrix.
lu = A.lu()
x = lu.solve(b) # Fast solve using cached LU factorization
Backends¶
Backend |
Device |
Description |
Recommended |
|---|---|---|---|
|
CPU |
SciPy backend using SuperLU or UMFPACK for direct solvers |
CPU default |
|
CPU |
Eigen C++ backend for iterative solvers (CG, BiCGStab) |
Alternative |
|
CUDA |
NVIDIA cuDSS for direct solvers (LU, Cholesky, LDLT) |
CUDA direct |
|
CUDA |
NVIDIA cuSOLVER for direct solvers |
Not recommended |
|
CUDA |
PyTorch-native iterative (CG, BiCGStab) with Jacobi preconditioning |
Large problems (>2M DOF) |
Methods¶
Direct Solvers¶
Method |
Backends |
Description |
Precision |
|---|---|---|---|
|
scipy |
SuperLU LU factorization (default for scipy) |
~1e-14 |
|
cudss, cusolver |
Cholesky factorization (for SPD matrices, fastest) |
~1e-14 |
|
cudss |
LDLT factorization (for symmetric matrices) |
~1e-14 |
|
cudss, cusolver |
LU factorization (general matrices) |
~1e-14 |
Iterative Solvers¶
Method |
Backends |
Description |
Precision |
|---|---|---|---|
|
scipy, eigen, pytorch |
Conjugate Gradient (for SPD) with Jacobi preconditioning |
~1e-6 |
|
scipy, eigen, pytorch |
BiCGStab (for general matrices) with Jacobi preconditioning |
~1e-6 |
|
scipy |
GMRES (for general matrices) |
~1e-6 |
Quick Start¶
Basic Usage¶
import torch
from torch_sla import SparseTensor
# Create a sparse matrix from dense (easier to read for small matrices)
dense = torch.tensor([[4.0, -1.0, 0.0],
[-1.0, 4.0, -1.0],
[ 0.0, -1.0, 4.0]], dtype=torch.float64)
# Create SparseTensor
A = SparseTensor.from_dense(dense)
# Solve Ax = b (auto-selects scipy+superlu on CPU)
b = torch.tensor([1.0, 2.0, 3.0], dtype=torch.float64)
x = A.solve(b)
CUDA Usage¶
import torch
from torch_sla import SparseTensor
# Create on CPU, move to CUDA (using the matrix from above)
A_cuda = A.cuda()
# Solve on CUDA (auto-selects cudss+cholesky for small problems)
b_cuda = b.cuda()
x = A_cuda.solve(b_cuda)
# For very large problems (DOF > 2M), use iterative
x = A_cuda.solve(b_cuda, backend='pytorch', method='cg')
Nonlinear Solve¶
Solve nonlinear equations with adjoint-based gradients:
from torch_sla import SparseTensor
# Create stiffness matrix
A = SparseTensor(val, row, col, (n, n))
# Define nonlinear residual: A @ u + u² = f
def residual(u, A, f):
return A @ u + u**2 - f
f = torch.randn(n, requires_grad=True)
u0 = torch.zeros(n)
# Solve with Newton-Raphson
u = A.nonlinear_solve(residual, u0, f, method='newton')
# Gradients flow via adjoint method
loss = u.sum()
loss.backward()
print(f.grad) # ∂L/∂f
Benchmark Results¶
2D Poisson equation (5-point stencil), NVIDIA H200 (140GB), float64:
Performance Comparison¶
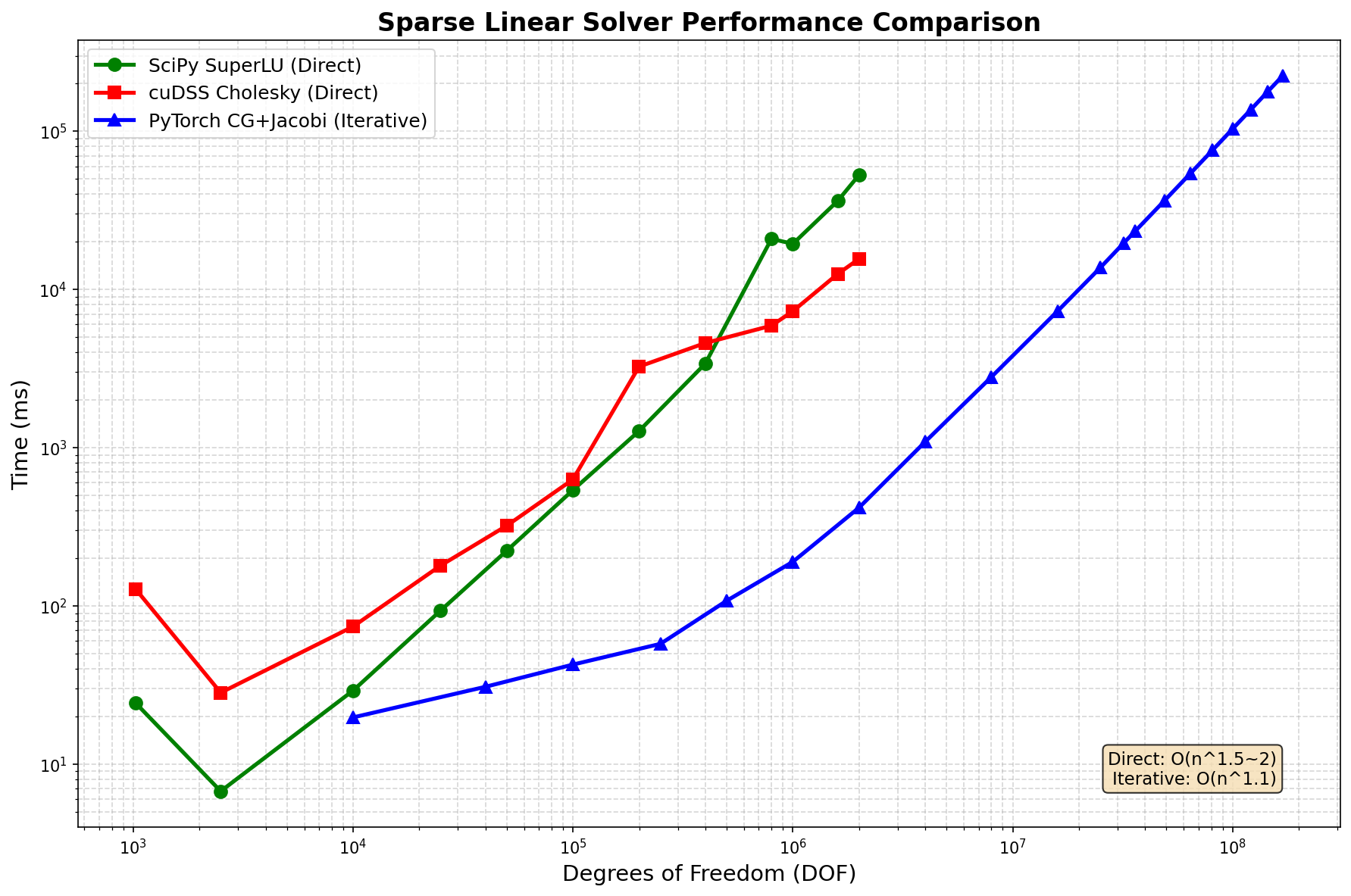
DOF |
SciPy SuperLU |
cuDSS Cholesky |
PyTorch CG+Jacobi |
Notes |
Winner |
|---|---|---|---|---|---|
10K |
24 |
128 |
20 |
All fast |
PyTorch |
100K |
29 |
630 |
43 |
SciPy |
|
1M |
19,400 |
7,300 |
190 |
PyTorch 100x |
|
2M |
52,900 |
15,600 |
418 |
PyTorch 100x |
|
16M |
OOM |
OOM |
7,300 |
PyTorch only |
|
81M |
OOM |
OOM |
75,900 |
PyTorch only |
|
169M |
OOM |
OOM |
224,000 |
PyTorch only |
Memory Usage¶
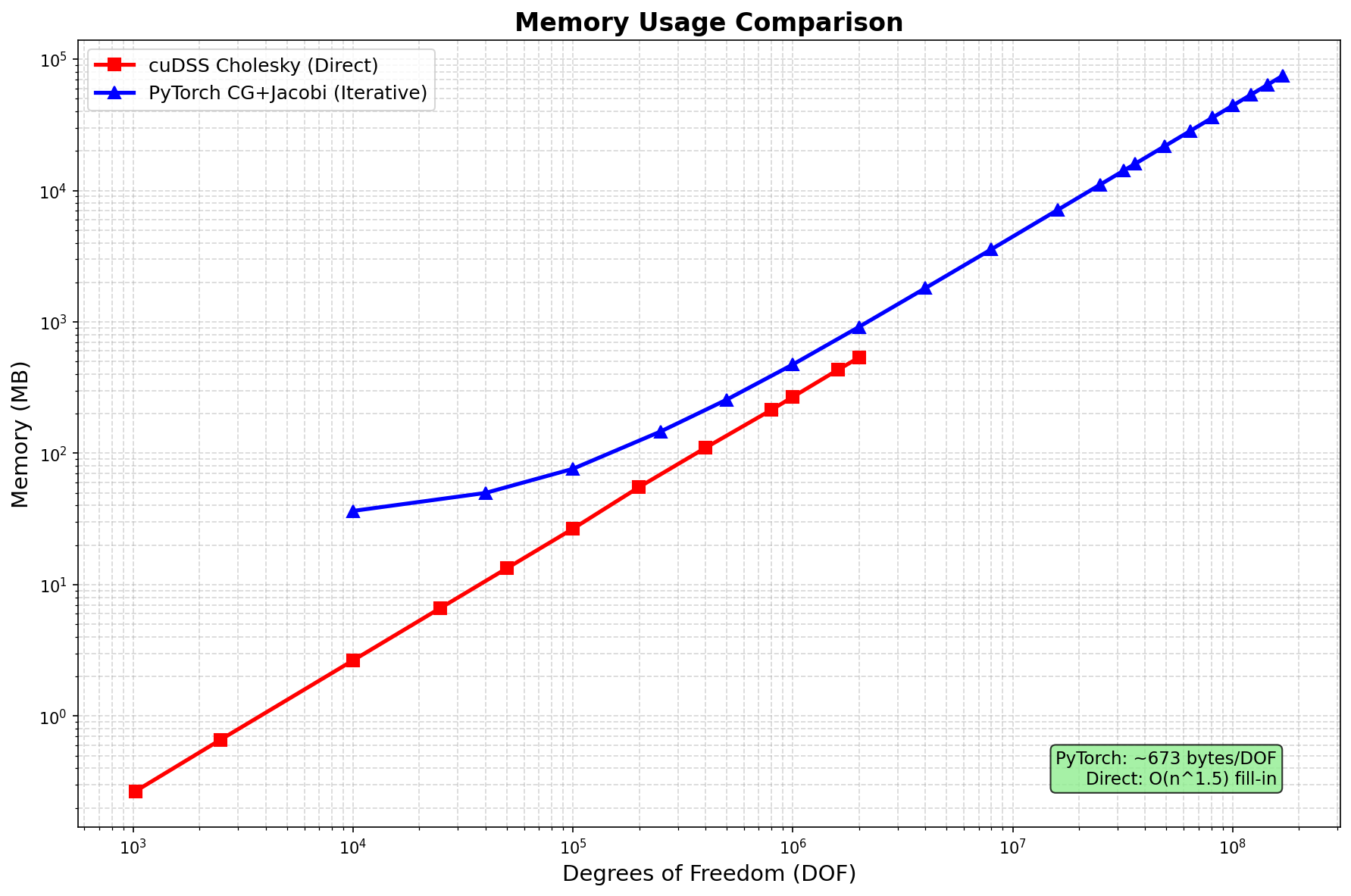
Method |
Memory Scaling |
Notes |
|---|---|---|
SciPy SuperLU |
O(n^1.5) fill-in |
CPU only, limited to ~2M DOF |
cuDSS Cholesky |
O(n^1.5) fill-in |
GPU, limited to ~2M DOF |
PyTorch CG+Jacobi |
O(n) ~443 bytes/DOF |
Scales to 169M+ DOF |
Accuracy¶
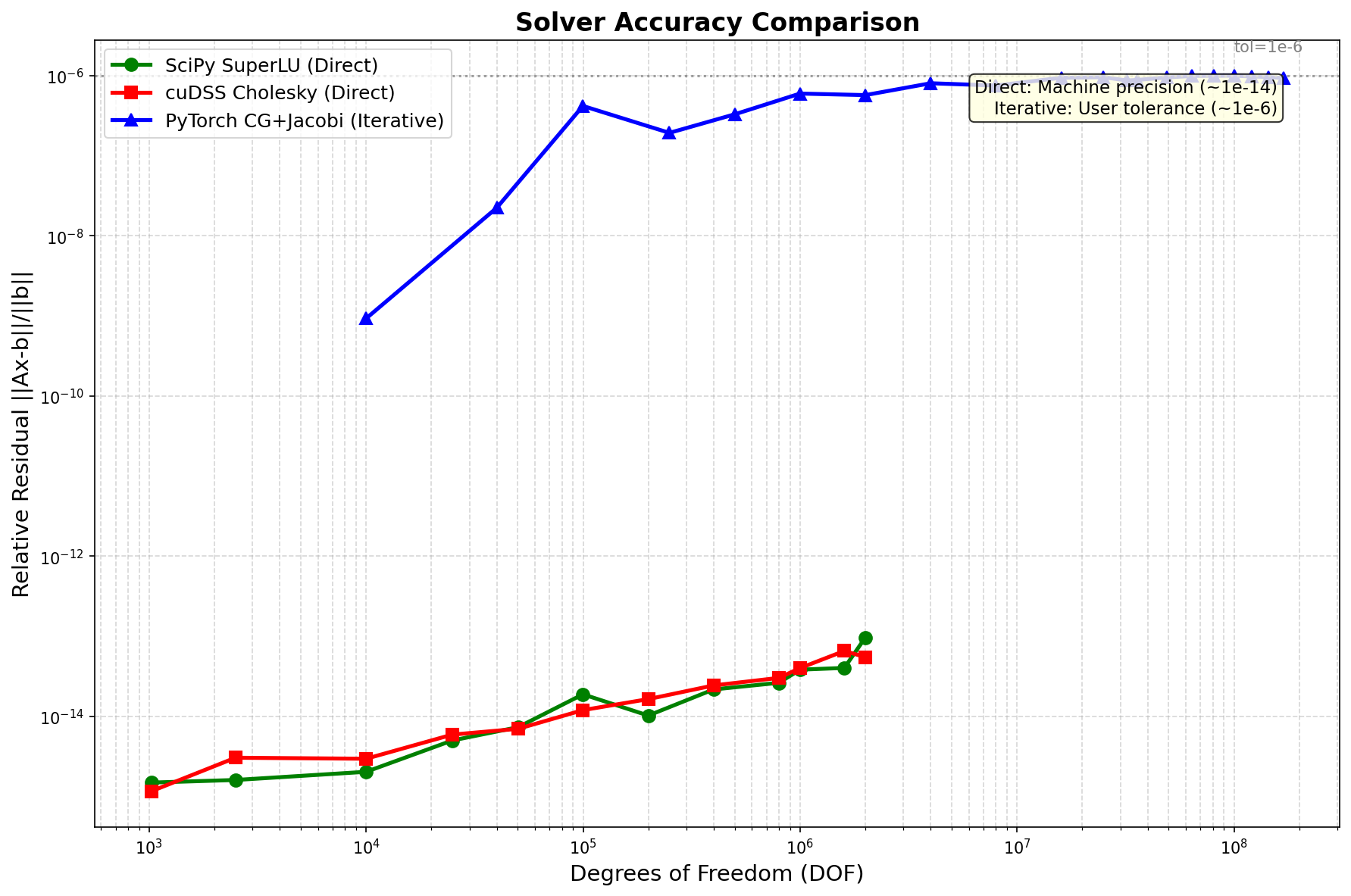
Method Type |
Relative Residual |
Notes |
|---|---|---|
Direct (scipy, cudss) |
~1e-14 |
Machine precision |
Iterative (pytorch+cg) |
~1e-6 |
User-configurable tolerance |
Key Findings¶
Iterative solver scales to 169M DOF with O(n^1.1) time complexity
Direct solvers limited to ~2M DOF due to O(n^1.5) memory fill-in
PyTorch CG+Jacobi is 100x faster than direct solvers at 2M DOF
Memory efficient: 443 bytes/DOF (vs theoretical minimum 144 bytes/DOF)
Trade-off: Direct solvers achieve machine precision, iterative achieves ~1e-6
Distributed Solve (Multi-GPU)¶
3-4x NVIDIA H200 GPUs with NCCL backend, scales to 400M+ DOF:
CUDA (3-4 GPU, NCCL):
DOF |
Time |
Memory/GPU |
GPUs |
|---|---|---|---|
10K |
0.1s |
0.03 GB |
4 |
100K |
0.3s |
0.05 GB |
4 |
1M |
0.9s |
0.27 GB |
4 |
10M |
3.4s |
2.35 GB |
4 |
50M |
15.2s |
11.6 GB |
4 |
100M |
36.1s |
23.3 GB |
4 |
200M |
119.8s |
53.7 GB |
3 |
300M |
217.4s |
80.5 GB |
3 |
400M |
330.9s |
110.3 GB |
3 |
Key Findings:
Scales to 400M DOF: Using 3x H200 GPUs (110 GB/GPU)
Near-linear scaling: 10M → 400M is 40x DOF, ~100x time
Memory efficient: ~275 bytes/DOF per GPU
CUDA 12x faster than CPU: 0.3s vs 7.4s for 100K DOF
# Run distributed solve with 3-4 GPUs
torchrun --standalone --nproc_per_node=3 examples/distributed/distributed_solve.py
Gradient Support¶
All operations support automatic differentiation via PyTorch autograd with O(1) graph nodes:
SparseTensor Gradient Support
Operation |
CPU |
CUDA |
Notes |
|---|---|---|---|
|
✓ |
✓ |
Adjoint method, O(1) graph nodes |
|
✓ |
✓ |
Adjoint method, O(1) graph nodes |
|
✓ |
✓ |
Power iteration, differentiable |
|
✓ |
✓ |
Adjoint, params only |
|
✓ |
✓ |
Standard autograd |
|
✓ |
✓ |
Sparse gradients |
|
✓ |
✓ |
Element-wise ops |
|
✓ |
✓ |
View-like, gradients flow through |
|
✓ |
✓ |
Standard autograd |
|
✓ |
✓ |
Standard autograd |
DSparseTensor Gradient Support
Operation |
CPU |
CUDA |
Notes |
|---|---|---|---|
|
✓ |
✓ |
Distributed matvec with gradient |
|
✓ |
✓ |
Distributed CG with gradient |
|
✓ |
✓ |
Distributed LOBPCG |
|
✓ |
✓ |
Distributed power iteration |
|
✓ |
✓ |
Distributed Newton-Krylov |
|
✓ |
✓ |
Distributed sum |
|
✓ |
✓ |
Gathers data (with warning) |
Key Features:
SparseTensor uses O(1) graph nodes via adjoint method for
solve(),eigsh()DSparseTensor uses true distributed algorithms (LOBPCG, CG, power iteration)
No data gather required for DSparseTensor core operations
For
nonlinear_solve(), gradients flow to the parameters passed toresidual_fn
Performance Tips¶
Use float64 for iterative solvers: Better convergence properties
Use cholesky for SPD matrices: 2x faster than LU
Use scipy+superlu for CPU: Best balance of speed and precision
Use cudss+cholesky for small CUDA problems: Fastest direct solver (< 2M DOF)
Use pytorch+cg for large problems: Memory efficient, scales to 169M+ DOF on single GPU
Use multi-GPU for very large problems: DSparseMatrix supports domain decomposition, 3 GPUs can reach 500M+ DOF
Avoid cuSOLVER: cudss is faster and supports float32
Use LU factorization for repeated solves: Cache with
A.lu()
Citation¶
If you use torch-sla in your research, please cite our paper:
Paper: arXiv:2601.13994 - Differentiable Sparse Linear Algebra with Adjoint Solvers and Sparse Tensor Parallelism for PyTorch
@article{chi2026torchsla,
title={torch-sla: Differentiable Sparse Linear Algebra with Adjoint Solvers and Sparse Tensor Parallelism for PyTorch},
author={Chi, Mingyuan},
journal={arXiv preprint arXiv:2601.13994},
year={2026},
url={https://arxiv.org/abs/2601.13994}
}